 
expression data using Cytoscape, Nature Protocols, 2, 2366-2382 (2007). The Cyni framework provides support for network inference through an extensible system of Cytoscape apps that provide key functionality in and around network inference, while eliminating the need for interaction with the much more complex Cytoscape application programmer interface (API Fig. Integration of biological networks and gene. found in protein interaction databases, including BioGRID and Pathwa圜ommons. A specic example of this workow: Cline, et al. The GeneMANIA Cytoscape app is capable of handling larger gene lists. Is this possible to do with Cytoscape? My aim is to actually see how gene-to-pathway information is structured on the network. 2.Load Attributes (Get data about networks). How do I overlay the pathway information to my genes on the network? I want to color code pathway information in such a way that different pathway will have different color. I attach the screenshot for PhosphoSitePlus and Wikipathways when I add info from Biogrid, Cytoscape continue working without give any errors or results. 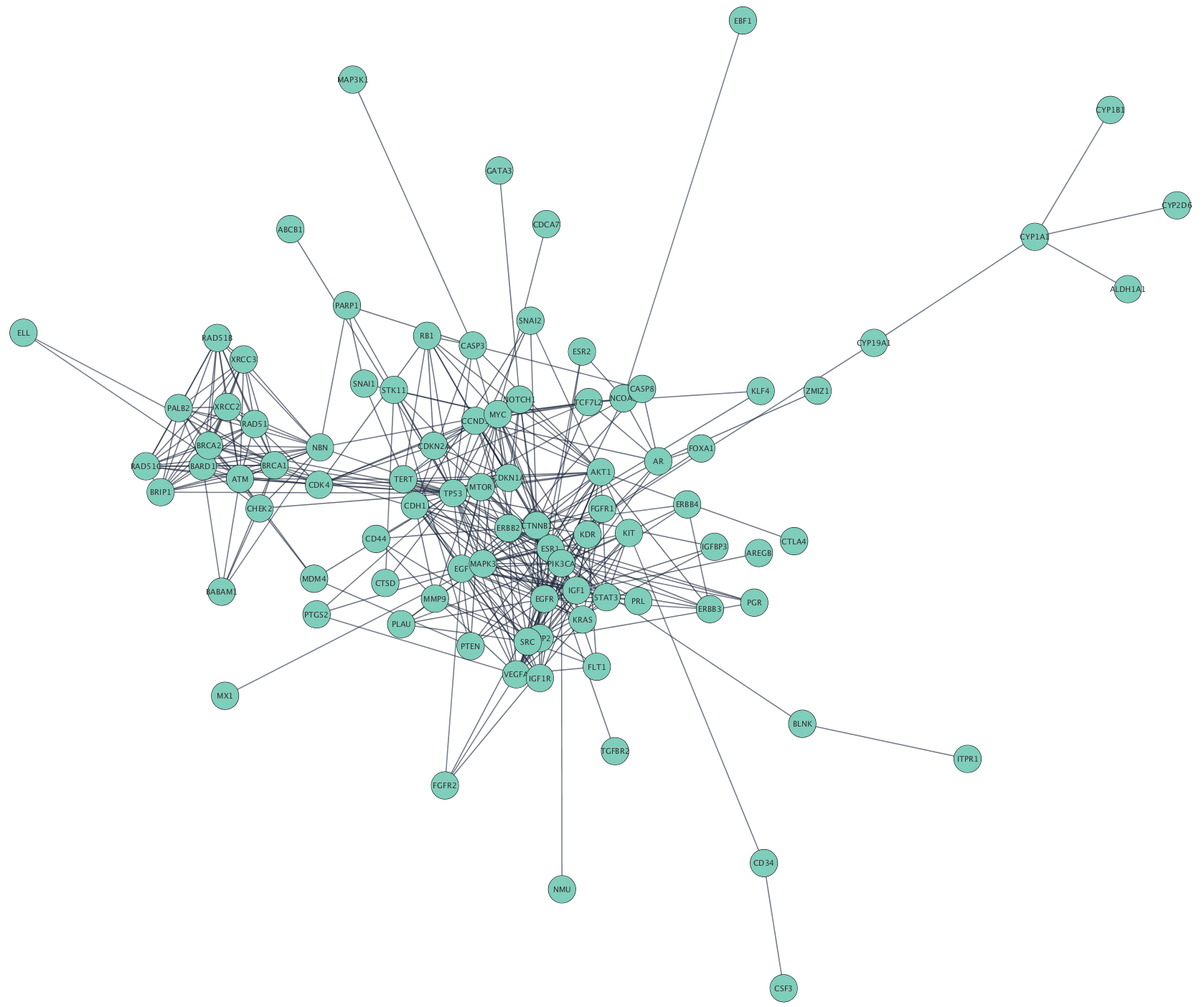
I'm using Cytoscape 3.2.1 which is the newest version. At the BioGRID, were focusing our curation efforts on the literature for these. The attribute file should like so considering one gene may map to multiple pathways gene1 pathway1 Cytoscape Single Cell Analysis Cytoscape Futures Introduction to Cytoscape and Network Biology Advanced Cytoscape Automation Intro to RC圓. I want to overlay pathway information to genes, mapped onto Biogrid database accessible via Cytoscape and visualize using Cytoscape.īasically I have about 200 overexpressed genes and I have obtained information about which pathways they map.Īccording to various answers here it seems I have to prepare two files - the overexpressed gene list file and the tab delimited attribute file( gene plus associated pathway) and import it into Cytoscape. 
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
